Laboratory of Processing and Analysis of Microscopic Images
Team

Head of lab: Anna Korzyńska, PhD, DSc, Assoc. Professor
Łukasz Roszkowiak, PhD Eng.,
Antonina Pater, MSc
Shrief Abdelazeez, MSc Eng.,
Krzysztof Siemion, MD,
Jakub Żak, MSc Eng.,
We are a small, interdisciplinary team developing innovative methods for analysing microscopic images of biological samples, contributing to diagnostics, prognosis, and biomedical research.
What We Do:
- Advanced Image Processing – We design methods for analysing stained tissue sections and cell smears, improving diagnostic precision and predictive modelling.
- Improving image analysis techniques using traditional and machine learning methods.
- Bridging Biology & Machine Learning – By integrating classical image analysis techniques with machine learning, we enhance image interpretation in medicine and life sciences.
Combining expertise from biology, computer science, and data analysis, we take a multidisciplinary approach to solving complex research challenges.
We also develop expertly annotated datasets, essential for advancing machine learning in biomedical applications. Some, like the Bialystok dataset (fragments from cervical cancer screening tests), are publicly available under link ibib.waw.pl/BOACD and described in the paper:
- Antonina Pater, Krzysztof Siemion, Karol Deptuch, Łukasz Roszkowiak, Jakub Żak, Katarzyna Jakubowska, Stanisław Sulkowski, Marek Baltaziak, Mariusz Koda, Anna Korzyńska (2023) "Conventional Cervical Cytology Image Dataset with Cell Outline Annotations", 2023 International Symposium on Image and Signal Processing and Analysis (ISPA), Rome, Italy, 2023, pp. 1-6, DOI: 10.1109/ISPA58351.2023.10279274
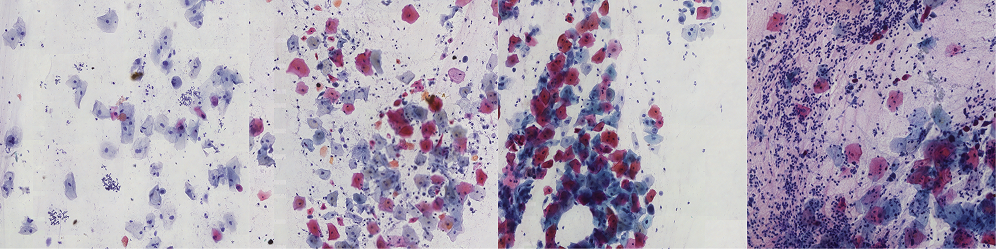
The Bialystok Dataset consists of 162 images (3500×3500 px) containing 2,419 annotated cervical cells, classified according to the Bethesda System. The images are extracted from whole slide images (WSI) of routine cervical smears, ensuring a diverse and realistic representation of real-world samples.
This dataset captures the full spectrum of cytological variability, ranging from clearly distinguishable cells to challenging cases—including stained mucus, densely packed dark cell clusters, and abundant neutrophils that obscure visibility. The dataset includes cytoplasm annotations categorized into six Bethesda classes and two additional groups: Unidentifiable cells and Unidentifiable cell clusters. These additional categories arise from the inherent uncertainty in cytodiagnostics, where some cells cannot be definitively classified but remain possible for segmentation and analysis.
Key Features:
- Realistic cervical smear fragments, reflecting routine cytological practice.
- Annotations aligned with the Bethesda System, ensuring clinical relevance.
- Versatility for segmentation, classification, and detection tasks.
- A challenging benchmark, designed to support the development of robust cytology AI models.
In the topic of Advanced Image Processing we have published following papers:
- Lukasz Roszkowiak, Anna Korzynska, Krzysztof Siemion, Jakub Zak, Dorota Pijanowska, Ramon Bosch, Marylene Lejeune & Carlos Lopez (2021) “System for quantitative evaluation of DAB&H-stained breast cancer biopsy digital images (CHISEL)”, Sci Rep 11, 9291 (2021). DOI: 10.1038/s41598-021-88611-y [IF 4.380, 140 points MNiSW].
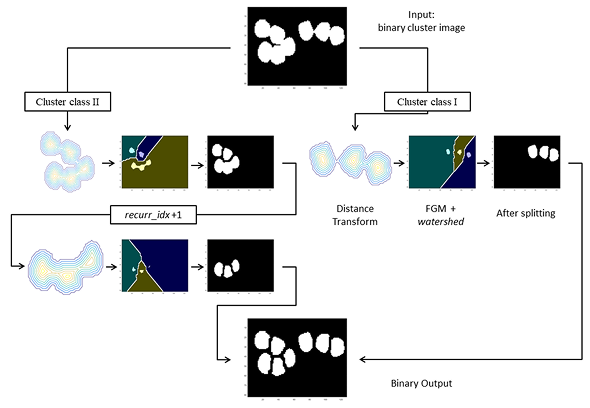
CHISEL (Computer-assisted Histopathological Image Segmentation and EvaLuation) is an end-to-end system designed for the quantitative analysis of digitized breast cancer biopsy samples stained with DAB&H. It features seamless segmentation through region of interest cropping, nuclei cluster splitting, and boundary refinement, utilizing machine learning and recursive local processing of distance transform to correct inaccurate outlines. Validated on labeled breast cancer datasets, CHISEL demonstrated reliable performance comparable to existing state-of-the-art methods. Its user-friendly interface is optimized for efficient computing, making it accessible to users with limited resources compared to those required by deep learning techniques.

- Antonina Pater, Łukasz Roszkowiak, Jakub Żak, Krzysztof Siemion, Anna Korzyńska, “An application of U-Net for cell detection in fragments of cytological smear images”, Proceedings of the 3rd Polish Conference on Artificial Intelligence, 2022, pp. 185-189, IBSN 978-83-7421-401-8
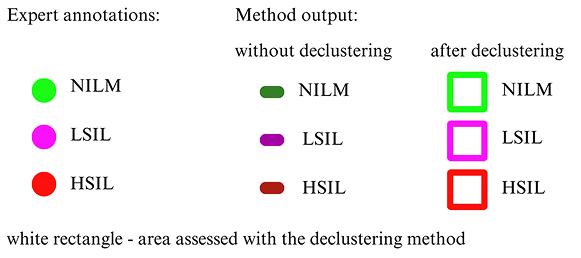
Cervical cytology is typically evaluated manually by identifying cells with precancerous changes. The Bethesda System for Reporting Cervical Cytology, a globally recognized reporting system, categorizes squamous intraepithelial lesions into two types: low-grade squamous intraepithelial lesion (LSIL) and high-grade squamous intraepithelial lesion (HSIL). Cells without abnormalities are classified as negative for intraepithelial lesions (NILM).
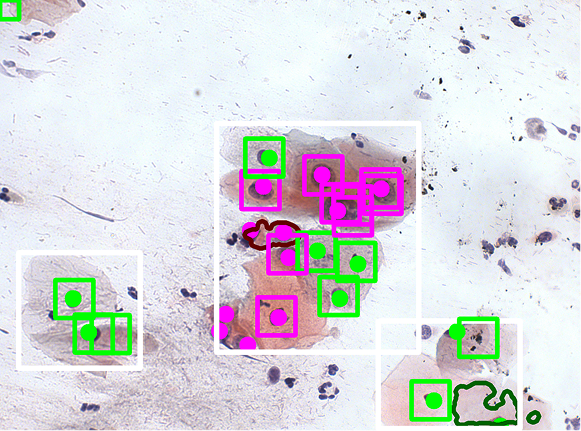
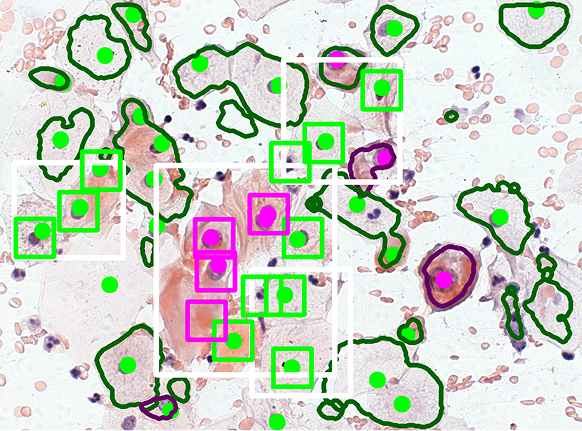
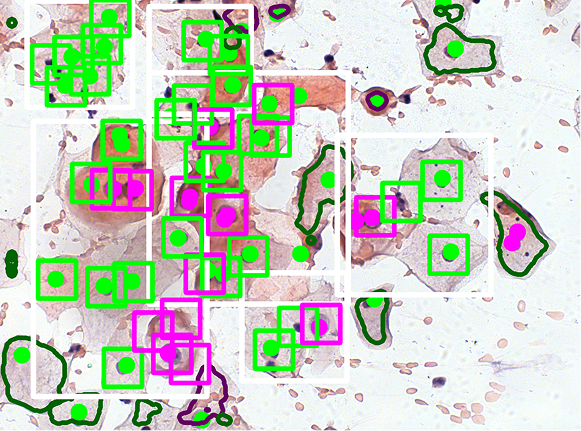
We have developed a method for classifying pre-segmented cells and nuclei using a VGG16-based model. To enhance the training process, we applied a GAN-based augmentation technique developed in our laboratory. The images below illustrate the classification results obtained after segmentation and the previously described declusterization process.


- 4. Antonina Pater, Łukasz Roszkowiak, Krzysztof Siemion, Anna Korzyńska (2021) „Estimation of the fraction of area covered by cells and cell clusters in WSI patches”. XXII Krajowa Konferencja Biocybernetyki i Inżynierii Biomedycznej, Warszawa, 19-21.05.2021.
- Jakub Żak, Krzysztof Siemion, Lukasz Roszkowiak, Anna Korzynska L (2021) „Fourier Transform Layer for Fast Forground Segmantation in Samples’ Images of Tissue Biopsies”) XXII National Conference on Biocybernetics and Biomedical Engineering, Warsaw, 19-21.05.2021.
- Krzysztof Siemion, Anna Korzyńska, Łukasz Roszkowiak, Jakub Żak, Antonina Pater, Joanna Reszeć-Giełażyn (2021) „Application of image analysis methods to evaluate histopathological slides in the study of prognostic factors of inflammatory spindle cell lesions” XXII National Biocybernetics and Biomedical Engineering Conference, Warsaw, 19-21.05.2021
- Łukasz Roszkowiak, Jakub Zak, Krzysztof Siemion, Antonina Pater, Anna Korzynska (2021) „Split point assessment for HRnet dual model” XXII National Biocybernetics and Biomedical Engineering Conference, Warsaw, 19-21.05.2021
In the topic of Improving image analysis techniques we have published following papers:
- Jakub Zak, Anna Korzynska, Antonina Pater, Lukasz Roszkowiak (2023) „Fourier Transform Layer: A proof of work in different training scenarios” Applied Soft Computing, 145, 110607. DOI: 10.1016/j.asoc.2023.110607 [IF 8.7; 200 points MNiSW].

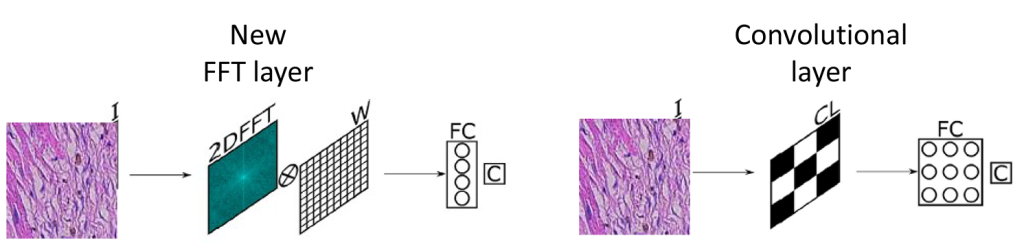
The paper presents a comparison of a new proposed approach that utilizes a connected neural network (NN) processing the Fourier Transform of images instead of the images themselves. Models consisting solely of this novel Fast Fourier Transform (FFT) layer were trained to classify images from publicly available datasets. The classification results were then compared with those of simpler models that included only one convolutional layer. Metrics such as test accuracy, Area Under the Curve (AUC), and training times were used for comparison. The findings indicated that the models employing the proposed FFT layer achieved test accuracies comparable to those of convolutional models—96% versus 98%—while demonstrating at least a 27% reduction in training time per epoch when run on a Central Processing Unit (CPU).
- Jakub Zak, Michal K. Grzeszczyk, Antonina Pater, Lukasz Roszkowiak, Krzysztof Siemion, Anna Korzynska (2022) „Cell image augmentation for classification task using GANs on Pap smear dataset”, Biocybernetics and Biomedical Engineering 42(3), pp 995-1011; DOI:10.1016/j.bbe.2022.07.003 [IF 5,667; 140 points MNiSW].

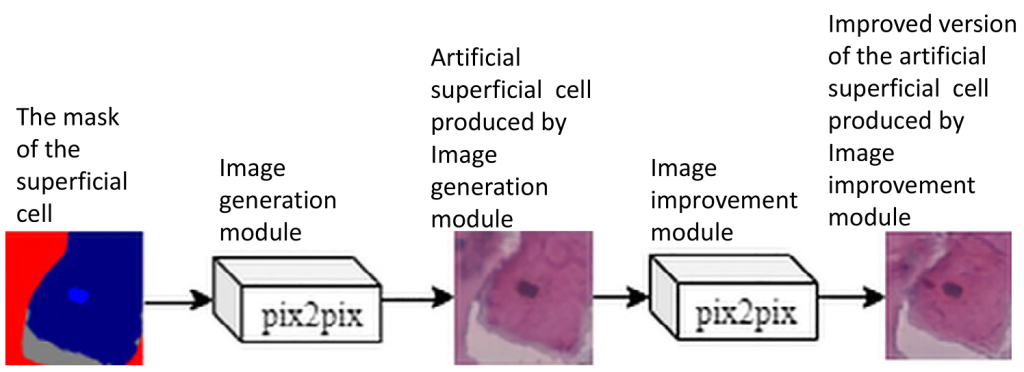
Generative Adversarial Networks (GANs) create artificial images that are challenging to distinguish from the real ones used during training. Incorporating these artificial images into the training process of Convolutional Neural Networks (CNNs) for detecting, segmenting, or classifying objects in images enhances model training and ultimately improves inference performance. This is especially crucial in the case of medical image analysis, which is subject to strict legal and ethical regulations. The paper proposes a two-step artificial image synthesis method based on the pix2pix network architecture. Validated on cervical smear cell classification dataset, the proposed method demonstrated boost in performance, particularly in classification accuracy.
- Anna Korzynska, Lukasz Roszkowiak, Jakub Zak, Krzysztof Siemion (2021) “A review of current systems for annotation of cell and tissue images in digital pathology” Biocybernetics and Biomedical Engineering vol 41, pp 1436-1453 [IF 4.314; 140 pts MNiSW].

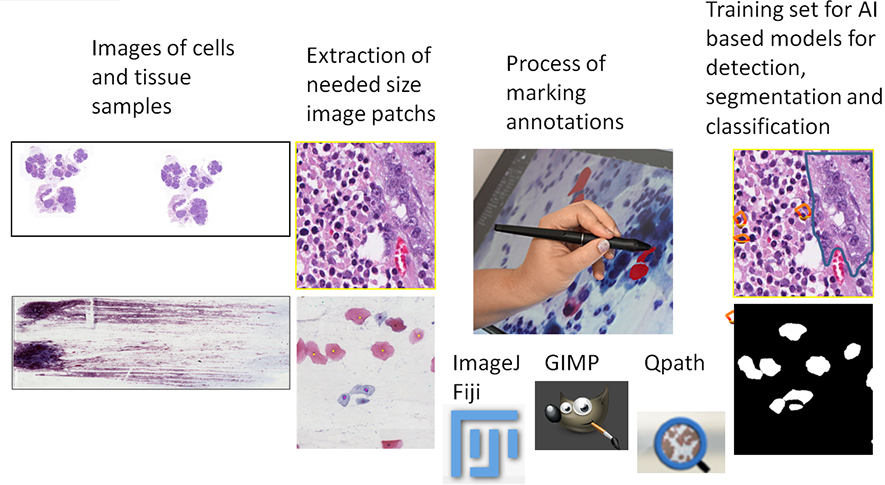
A significant limitation in computer-assisted diagnostic systems for pathology is the high cost of data annotation. Evaluating tissue (histopathological) and cellular (cytological) specimens presents a complex challenge. To streamline the labor-intensive process of collecting a sufficiently large dataset, various systems: GIMP, QuPath, ImageJ can be utilized for image annotation.
- Krzysztof Siemion, Lukasz Roszkowiak, Jakub Zak, Antonina Pater, Anna Korzynska (2024) Tissue Pattern Classification with CNN in Histological Images. In: Strumiłło, P., Klepaczko, A., Strzelecki, M., Bociąga, D. (eds) The Latest Developments and Challenges in Biomedical Engineering. PCBEE 2023 Lecture Notes in Networks and Systems, vol 746. Springer, Cham. DOI: 10.1007/978-3-031-38430-1_2
- Shrief Abdelazeez, Lukasz Roszkowiak, Krzysztof Siemion, Carlos Lopez, Marylene Lejeune, Anna Korzynska (2023) „T regulatory Lymphocyte Nuclei Segmentation in DAB - Stained Breast Cancer Biopsy Digital Images Using Deep Learning Algorithms”; in Book of Abstracts - Polish Conference on Biocybernetics and Biomedical Engineering (PCBEE) Ed. Paweł Strumiłło, Artur Klepaczko, Michał Strzelecki, Bociąga, pp 41
- Jakub Zak, Krzysztof Siemion, Lukasz Roszkowiak, Anna Korzynska (2022) „Fourier Transform Layer for Fast Forground Segmantation in Samples’ Images of Tissue Biopsies”, in Biocybernetics and Biomedical Engineering - Current Trends and Challenges Ed. D. Pijanowska, K. Zielinski, A. Liebert, J Kacprzyk; Lecture Notes in Networks and Systems 293, pp 118-125, Springer 2022 DOI: 10.1007/978-3-030-83704-4
In the topic of Bridging Biology & Machine Learning we have published following papers:
- Paweł Pstrokoński, Lukasz Roszkowiak, Anna Korzyńska, Wojeciech Wójcik, Martin Packert, Joanna Rosenberger, Dominika Mierzwa-Szymkowiak, Magdalena Seplowska, Jan Lontkowski, Marek Słupek, Krzysztof Damaziak (2025) “Can Explainable AI classify shrike (Laniidae) eggs by uncovering species-wide pigmentation patterns?” PLOS ONE accepted, [IF 2.9; 100 p. MNiSW].

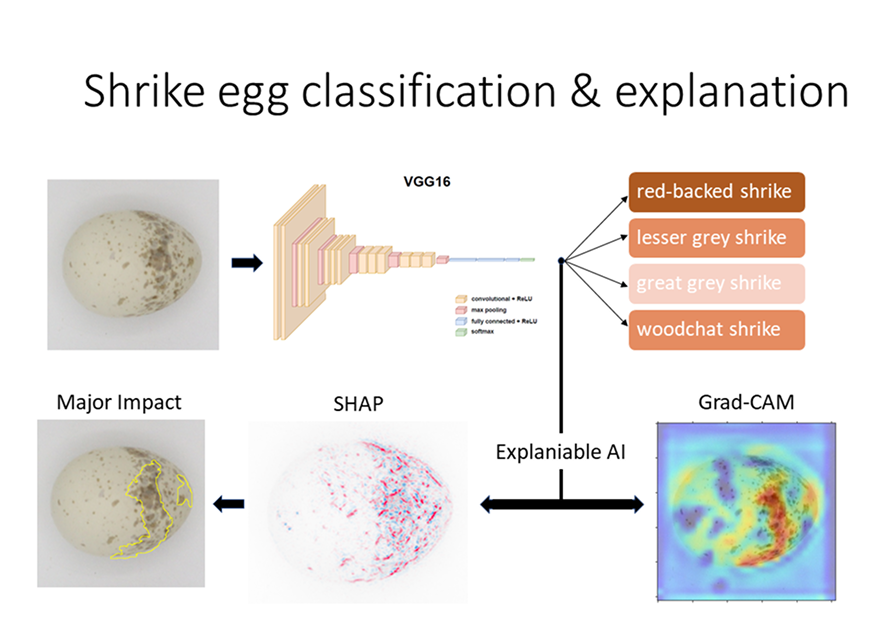
Can Explainable AI classify shrike (Laniidae) eggs by uncovering species-wide pigmentation patterns?
Computer analysis of egg pigmentation can reveal identification patterns for bird species, which will contribute to the verification of research hypotheses. The research is conducted on photographed archival specimens of eggs (eggshells) belonging to one genus Lanius with its 5 species. A systematic analysis of classification performance using the deep convolutional network VGG16 was performed. The impact of different image augmentation strategies was compared and, using the knowledge gained, the dataset was rebalanced, finally achieving an accuracy above 95% on the test set.
This study applies Explainable AI to analyze pigmentation patterns on bird eggs, focusing on the genus Lanius, which exhibits high intra-species variability. Using CNNs, the research achieved over 95% accuracy in species classification. To uncover species-wide identification signatures, Grad-CAM and SHAP deep explainers were applied, though they primarily highlighted general trends rather than specific discriminative features. This method enhances museum collection organization, species verification, and oological study accuracy while also offering insights into avian evolution, ecological strategies, and potential biomedical applications.

EGGS is a specialized image processing approach designed to create a standardized and high-quality dataset of avian egg images. It integrates advanced preprocessing, orientation detection, and damaged specimen removal to improve oological data analysis. By combining these techniques with deep learning models, EGGS enhances ecological and heritage research, offering a precise framework for egg annotation. The dataset created with EGGS serves as a valuable resource for ornithological studies, enabling deeper insights into avian reproduction, evolution, and environmental adaptation.
- Łukasz Roszkowiak, Paweł Pstrokoński, Wojeciech Wójcik, Krzysztof Damaziak, Anna Korzyńska (2025) „EGGS: Efficient Gathering and Structuring of Avian Egg Datasets”, 24th Polish Conference on Biocybernetics and Biomedical Engineering conference, Warsaw, 16-18.06.2025
The additional publications:
In cooperation with Medical University of Bialystok:
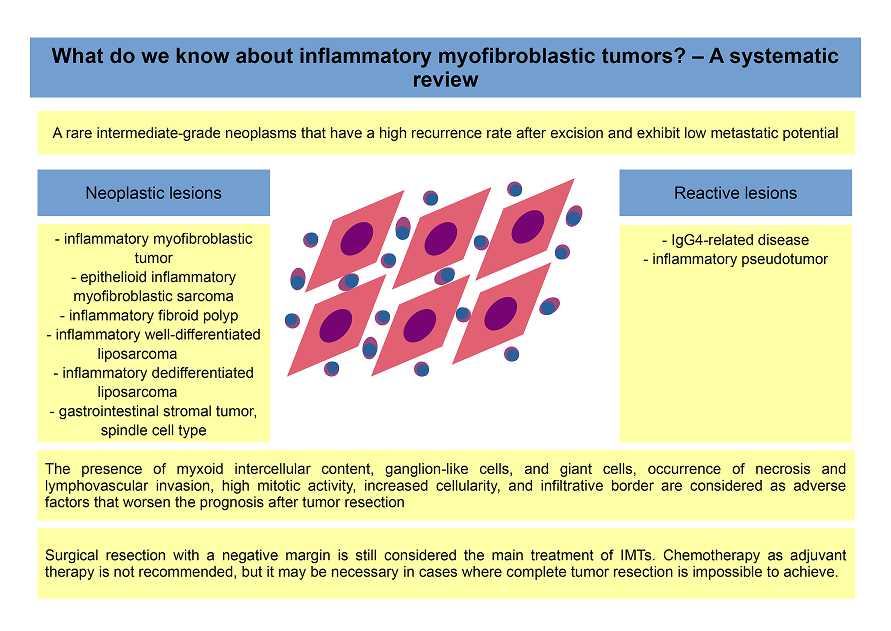
- Krzysztof Siemion, Joanna Reszec-Gielazyn, Joanna Kisluk, Lukasz Roszkowiak, Jakub Zak, Anna Korzynska: „What do we know about inflammatory myofibroblastic tumors? - A systematic review” Advances in Medical Sciences 67 (1), 2022 pp.129-138 DOI: 0.1016/j.advms.2022.02.002, [IF 2,852, 100pt ].

This systematic review explores the current understanding of inflammatory myofibroblastic tumors (IMTs), rare mesenchymal neoplasms with variable clinical behavior. It summarizes key findings related to their pathogenesis, histopathological features, diagnostic challenges, and treatment strategies. The review also highlights recent advances in molecular genetics, particularly ALK gene rearrangements, which have important diagnostic and therapeutic implications.
In cooperation with Research in Oncological Pathology and Bioinformatics (PO&B) (formerly known as Molecular Biology and Research Section), Institut d’Investigacio Sanitaria Pere Virgili (IISPV), URV, Spain:
- Marylène Lejeune, Benoît Plancoulaine, Nicolas Elie, Ramon Bosch, Laia Fontoura, Izar de Villasante, Anna Korzyńska; at all (2021) “How the variability between computer-assisted analysis procedures evaluating immune markers can influence patients’ outcome prediction”.”; Histochemistry and Cell Biology; DOI: 10.1007/s00418-021-02022-8 [IF 4.304, 100pts ]
- Carlos López, Albert Gibert-Ramos, Ramón Bosch, Anna Korzynska, Marcial García-Rojo, Gloria Bueno, Joan Francesc García-Fontgivell, Salomé Martínez González, Laia Fontoura, Andrea Gras Navarro, Esther Sauras Colón, Júlia Casanova Ribes, Lukasz Roszkowiak, Albert Roso, Marta Berenguer, Montserrat Llobera, Jordi Baucells, Marylène Lejeune (2021) “Differences in the Immune Response of the Nonmetastatic Axillary Lymph Nodes between Triple-Negative and Luminal A Breast Cancer Surrogate Subtypes” American Journal of Pathology (2021) Volume 191, Issue 3, March 2021, Pages 545-554 DOI: 10.1016/j.ajpath.2020.11.008 [IF 4.307, 140 points MNiSW].
- Carlos López, Ramón Bosch, Anna Korzynska,-Marcial García-Rojo, Gloria Bueno, Joan Francesc García Fontgivell, Salomé Martínez González, Andrea Gras Navarro, Esther Sauras Colón, Júlia Casanova Ribes, Lukasz Roszkowiak, Daniel Mata, Meritxell Arenas, Junior Gómez, Albert Roso, Marta Berenguer, Silvia Reverté Villarroya, Montserrat Llobera, Jordi Baucells, Marylène Lejeune () „CD68 and CD83 immune populations in non metastatic axillary lymph nodes are of prognostic value for the survival and relapse of breast cancer patients”, Breast Cancer 29(4) pp.618-635 DOI: 10.1007/s12282-022-01336-2, [IF 3.303; 70 points MNiSW].
- Marylène Lejeune, Laia Reverté, Esther Sauras, Noèlia Gallardo, Ramon Bosch, Albert Roso, Anna Petit, Vicente Peg, Francisco Riu, Joan García-Fontgivell, José Ibáñez, Fernanda Relea,Begoña Vieites, Catherine Bor, Luis de la Cruz-Merino, Meritxell Arenas, Valerie Rodriguez, Juana Galera, Anna Korzynska, Philippe Belhomme, Benoît Plancoulaine, Tomás Álvaro and Carlos López (2023) „Prognostic Implications of the Residual Tumor Microenvironment after Neoadjuvant Chemotherapy in Triple-Negative Breast Cancer Patients without Pathological Complete Response”, Cancers 2023, 15, 597. DOI: 10.3390/cancers15030597 [IF 5.2; 200 points MNiSW].
- Marylène Lejeune, Laia Reverté, Noèlia Gallardo, Esther Sauras, Ramon Bosch 1 , Daniel Mata, Albert Roso, Anna Petit, Vicente Peg, Francisco Riu, Joan García-Fontgivell, Fernanda Relea, Begoña Vieites, Luis de la Cruz-Merino, Meritxell Arenas, Valeri Rodriguez, Juana Galera, Anna Korzynska, Benoît Plancoulaine, Tomás Álvaro and Carlos López 92023) „Matrix Metalloproteinase-9 Expression Is Associated with the Absence of Response to Neoadjuvant Chemotherapy in Triple-Negative Breast Cancer Patients”, International Journal of Molecular Sciences 24, 11297. DOI: 10.3390/ijms241411297 [IF 5.6 ;140 points MNiSW] Cooperation in consortium developing project BosomShield:
- Alessio Fiorin, Laia Adalid Llansa, Elena Goyda, Vincenzo Della Mea, Anna Korzynska, Shrief Abdelazeez, Ramon Bosch Príncep, Alba Fischer Carles, Noelia Gallardo Borràs, Marylène Lejeune, Daniel Mata Cano, Domenec Puig, Hatem A. Rashwan, Esther Sauras Colón, Mikel Relloso Ortiz de Uriarte, Laia Reverté Calvet, Carlos López Pablo (2024). Optimising Region of Interest Registration for Multiple-Tissue Whole Slide Images. In: Modat, M., Simpson, I., Špiclin, Ž., Bastiaansen, W., Hering, A., Mok, T.C.W. (eds) Biomedical Image Registration. WBIR 2024 Lecture Notes in Computer Science, vol 15249. springer, Cham. DOI: 10.1007/978-3-031-73480-9_26
Keywords:
image analysis, image segmentation, microscopic images, time laps microscopy, analysis of WSI, correction and standardization of scanned images of stained tissues, digital pathology, computer-aided quantitative morphometry, breast cancer investigation, cervical smear images analysis, avian biology, oology
The Laboratory has concluded following projects:
- “A comprehensive CAD system based on radiologic- and pathologic-image biomarkers for diagnosis and prognosis of breast cancer relapse , HORIZON-MSCA-2021-DN-01-01 MSCA Doctoral Network Project No. 10107322 acronym BosomShield; started October 2022.
- “The segmentation method for cell segmentation in sequence of microscopic images”.” financed by National Sciences Center (NCN), headed by dr Anna Korzyńska, developed from 10.08.2008 to 09.06.2011;
- “Development of automatic quantitative methods for immunohistochemicaly stained thin tissue sections in investigation of immune response in breast cancer” financed by Institute of Health Carlos III in Spain, headed by dr Carlos Lopez from Molecular Biology and Research Section, Hospital Verge de la Cinta w Tortozie, developed from 1.01.2013 to 31.06.2016,
- “Automated analysis of tumour microenvironment in triple negative breast cancer without complete pathological response to neoadjuvant chemotherapy. Predictive factor of relapse” financed by Institute of Health Carlos III in Spain, headed by dr Merylene Lejeune from Molecular Biology and Research Section, Hospital Verge de la Cinta w Tortozie, developed from 1.01.2014 to 31.12.2016,
- „Web-based platform for the computer analysis of microscopic images to support the pathological diagnosis”.” financed by Institute of Health Carlos III in Spain, headed by dr Merylene Lejeune from Molecular Biology and Research Section, Hospital Verge de la Cinta w Tortozie, developed from 1.01.2014 to 31.12.2016,
- "Supporting software to assist pathologist in evaluation of immunohistochemically stained tissue samples of breast cancer stained with DAB&H” financed by National Sciences Center (NCN), headed by Łukasz Roszkowiak developed from 2014 to 2019.
Defended doctoral thesis:
- “New method of computer aided quantification of cells in whole-slide images of immunohistochemically stained breast cancer tissue samples”, Łukasz Roszkowiak – defended with honors in 2023.
In 2023, Lukasz Roszkowiak has achieved the academic degree of Doctor (PhD) in the field of engineering and technical sciences in the discipline of biomedical engineering, after presentation of doctoral dissertation entitled “New method of computer aided quantification of cells in whole-slide images of immunohistochemically stained breast cancer tissue samples”. This dissertation focused on methods for nuclei quantification, segmentation, and classification, incorporating adaptive thresholding, cluster splitting, and AI-based processing to enhance accuracy and efficiency. A key achievement was CHISEL, an end-to-end system designed for precise and repeatable cell analysis, improving digital pathology workflows. Beyond CHISEL, the research explored interpolation methods for virtual slides and strategies to minimize object loss, contributing to more reliable diagnostic and biomedical imaging techniques. Assoc. Prof. Anna Korzyńska, PhD, DSc was the supervisor.
Equipment:
In the Laboratory, there is available an optical microscope, working in bright field or fluorescence modes. It has capabilities of digitally recording sequences with PC-controlled irises (for UV and visible light) and microscope table in Z axis. A high-performance computing station equipped with an NVIDIA RTX A5000 GPU, a high-end Intel Pentium processor, and 128 GB of RAM, designed for demanding computational and image-oriented tasks.